elektronn2.neuromancer package¶
Submodules¶
elektronn2.neuromancer.computations module¶
-
elektronn2.neuromancer.computations.apply_activation(x, activation_func, b1=None)[source]¶ Return an activation function callable matching the name Allowed names: ‘relu’, ‘tanh’,’prelu’, ‘sigmoid’, ‘maxout <i>’, ‘lin’,’abs’,’soft+’, ‘elu’, ‘selu’.
Parameters: - x (T.Tensor) – Input tensor.
- activation_func (str) – Name of the activation function.
- b1 – Optional b1 parameter for the activation function. If this is None, no parameter is passed.
Returns: Activation function applied to
x.Return type: T.Tensor
-
elektronn2.neuromancer.computations.apply_except_axis(x, axis, func)[source]¶ Apply a contraction function on all but one axis.
Parameters: - x (T.Tensor) – Input tensor.
- axis (int) – Axis to exclude on application.
- func (function) – A function with signature
func(x, axis=)eg T.mean, T.std …
Returns: Contraction of
x, but of the same dimensionality.Return type: T.Tensor
-
elektronn2.neuromancer.computations.conv(x, w, axis_order=None, conv_dim=None, x_shape=None, w_shape=None, border_mode='valid', stride=None)[source]¶ Apply appropriate convolution depending on input and filter dimensionality. If input
w_shapeis known, conv might be replaced by tensordotThere are static assumptions which axes are spatial.
Parameters: - x (T.Tensor) – Input data (mini-batch).Tensor of shape
(b, f, x),(b, f, x, y),(b, z, f, x, y)or(b,f,x,y,z). - w (T.Tensor) – Set of convolution filter weights.Tensor of shape
(f_out, f_in, x),(f_out, f_in, x, y),(f_out, z, f_in, x, y)or(f_out, f_in, x, y, z). - axis_order (str) – (only relevant for 3d)
'dnn'(b,f,x,y(,z))or'theano'(b, z, f, x, y). - conv_dim (int) – Dimensionality of the applied convolution (not the absolute dim of the inputs).
- x_shape (tuple) – shape tuple (
TaggedShapesupported). - w_shape (tuple) – shape tuple, see
w. - border_mode (str) –
'valid': only apply filter to complete patches of the image. Generates output of shape: image_shape -filter_shape + 1.'full'zero-pads image to multiple of filter shape to generate output of shape: image_shape + filter_shape - 1.
- stride (tuple) – (tuple of len 2)Factor by which to subsample the output.
Returns: Set of feature maps generated by convolution.
Return type: T.Tensor
- x (T.Tensor) –
-
elektronn2.neuromancer.computations.dot(x, W, axis=1)[source]¶ Calculate a tensordot between 1 axis of
xand the first axis ofW.Requires
x.shape[axis]==W.shape[0]. Identical to dot ifx,W2d andaxis==1.Parameters: - x (T.Tensor) – Input tensor.
- W (T.Tensor) – Weight tensor, (f_in, f_out).
- axis (int) – Axis on
xto apply dot.
Returns: xwith dot applied. The shape ofxchanges onaxiston_out.Return type: T.Tensor
-
elektronn2.neuromancer.computations.fragmentpool(conv_out, pool, offsets, strides, spatial_axes, mode='max')[source]¶
-
elektronn2.neuromancer.computations.fragments2dense(fragments, offsets, strides, spatial_axes)[source]¶
-
elektronn2.neuromancer.computations.maxout(x, factor=2, axis=None)[source]¶ Maxpooling along the feature axis.
The feature count is reduces by
factor.Parameters: - x (T.Tensor) – Input tensor (b, f, x, y), (b, z, f, x, y).
- factor (int) – Pooling factor.
- axis (int or None) – Feature axis of
x(1 or 2). If None, 5d tensors get axis 2 and all others axis 1.
Returns: xwith pooling applied.Return type: T.Tensor
-
elektronn2.neuromancer.computations.pooling(x, pool, spatial_axes, mode='max', stride=None)[source]¶ Pooling along spatial axes of 3d and 2d tensors.
There are static assumptions which axes are spatial. The spatial axes must be divisible by the corresponding pooling factor, otherwise the computation might crash later.
Parameters: - x (T.Tensor) – Input tensor (b, f, x, y), (b, z, f, x, y).
- pool (tuple) – 2/3-tuple of pooling factors.
They refer to the spatial axes of
x(x,y)/(z,x,y). - spatial_axes (tuple) –
- mode (str) –
Can be any of the modes supported by Theano’s dnn_pool(): (‘max’, ‘average_inc_pad’, ‘average_exc_pad’, ‘sum’).
’max’ (default): max-pooling ‘average’ or ‘average_inc_pad’: average-pooling ‘sum’: sum-pooling
- stride (tuple) –
Returns: xwith maxpooling applied. The spatial axes are decreased by the corresponding pooling factorsReturn type: T.Tensor
-
elektronn2.neuromancer.computations.softmax(x, axis=1, force_builtin=False)[source]¶ Calculate softmax (pseudo probabilities).
Parameters: - x (T.Tensor) – Input tensor.
- axis (int) – Axis on which to apply softmax.
- force_builtin (bool) – force usage of
theano.tensor.nnet.softmax(more stable).
Returns: xwith softmax applied, same shape.Return type: T.Tensor
-
elektronn2.neuromancer.computations.unpooling(x, pool, spatial_axes)[source]¶ Symmetric unpooling with border: s_new = s*pool + pool-1.
Insert values strided, e.g for pool=3: 00x00x00x…00x00.
Parameters: - x (T.Tensor) – Input tensor.
- pool (int) – Unpooling factor.
- spatial_axes (list) – List of axes on which to perform unpooling.
Returns: xwith unpooling applied.Return type: T.Tensor
-
elektronn2.neuromancer.computations.upconv(x, w, stride, x_shape=None, w_shape=None, axis_order='dnn')[source]¶
-
elektronn2.neuromancer.computations.upsampling(x, pool, spatial_axes)[source]¶ Upsamling through repetition: s_new = s*p.
e.g for pool=3: aaabbbccc…
Parameters: - x (T.Tensor) – Input tensor.
- pool (int) – Upsampling factor.
- spatial_axes (list) – List of axes on which to perform upsampling.
Returns: xwith upsampling applied.Return type: T.Tensor
elektronn2.neuromancer.graphmanager module¶
elektronn2.neuromancer.graphutils module¶
-
class
elektronn2.neuromancer.graphutils.TaggedShape(shape, tags, strides=None, mfp_offsets=None, fov=None)[source]¶ Bases:
objectObject to manage shape and associated tags uniformly. The
[]-operator can be used get shape values by either index (int) or tag (str)Parameters: - shape (list/tuple of int) – shape of array, unspecified shapes are
None - tags (list/tuple of strings or comma-separated string) – tags indicate which purpose the dimensions of the tensor serve. They are sometimes used to decide about reshapes. The maximal tensor has tags: “r, b, s, f, z, y, x, s” which denote: * r: perform recurrence along this axis * b: batch size * s: samples of the same instance (over which expectations are calculated) * f: features, filters, channels * z: convolution no. 3 (slower than 1,2) * y: convolution no. 1 * x: convolution no. 2
- strides – list of strides, only for spatial dimensions, so it is 1-3 long
- mfp_offsets –
-
addaxis(axis, size, tag)[source]¶ Create new TaggedShape with new axis inserted at
axisof sizesizetaggedtag. If axis is a tag, the new axis is right of that tag
-
delaxis(axis)[source]¶ Create new TaggedShape with new axis inserted at
axisof sizesizetaggedtag. If axis is a tag, the new axis is right of that tag
-
ext_repr¶
-
fov¶
-
fov_all_centered¶
-
mfp_offsets¶
-
ndim¶
-
offsets¶
-
shape¶
-
spatial_axes¶
-
spatial_shape¶
-
spatial_size¶
-
strides¶
-
stripbatch_prod¶ Calculate product excluding batch dimension
-
stripnone¶ Return the shape but with all None elements removed (e.g. if batch size is unspecified)
-
stripnone_prod¶ Return the product of the shape but with all None elements removed (e.g. if batch size is unspecified)
-
updatefov(axis, new_fov)[source]¶ Create new TaggedShape with
new_fovonaxis. Axis is given as index of the spatial axes (not matching the absolute index of sh).
- shape (list/tuple of int) – shape of array, unspecified shapes are
-
class
elektronn2.neuromancer.graphutils.make_func(tt_input, tt_output, updates=None, name='Unnamed Function', borrow_inp=False, borrow_out=False, profile_execution=False)[source]¶ Bases:
objectWrapper for compiled theano functions. Features:
- The function is compiled on demand (i.e. no wait at initialisation)
- Singleton return values are returned directly, multiple values as list
- The last execution time can inspected in the attribute
last_exec_time - Functions can be timed:
profile_executionis anintthat specifies the number of runs to average. The average time is printed then. - In/Out values can have a
borrowflag which might overwrite the numpy arrays but might speed up execution (see theano doc)
elektronn2.neuromancer.loss module¶
-
class
elektronn2.neuromancer.loss.GaussianNLL(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeSimilar to squared loss but “modulated” in scale by the variance.
Parameters: Computes element-wise:
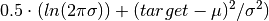
-
class
elektronn2.neuromancer.loss.BinaryNLL(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeBinary NLL node. Identical to cross entropy.
Parameters: Computes element-wise:
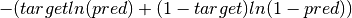
-
class
elektronn2.neuromancer.loss.AggregateLoss(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeThis node is used to average the individual losses over a batch (and possibly, spatial/temporal dimensions). Several losses can be mixed for multi-target training.
Parameters: - parent_nodes (list/tuple of graph or single node) – each component is some (possibly element-wise) loss array
- mixing_weights (list/None) – Weights for the individual costs. If none, then all are weighted
equally. If mixing weights are used, they can be changed during
training by manipulating the attribute
params['mixing_weights']. - name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
- The following is all wrong, mixing_weights are directly used (#) –
- losses are first summed per component, and then the component sums (The) –
- summed using the relative weights. The resulting scalar is finally (are) –
- such that (normalised) –
- The cost does not grow with the number of mixed components
- Components which consist of more individual losses have more weight e.g. If there is a constraint on some hidden representation with 20 features and a constraint the reconstruction of 100 features, the reconstruction constraint has 5x more impact on the overall loss than the constraint on the hidden state (provided those two loss are initially on the same scale). If they are intended to have equal impact, the weights should be used to upscale the constraint against the reconstruction.
-
class
elektronn2.neuromancer.loss.SquaredLoss(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeSquared loss node.
Parameters: - pred (Node) – Prediction node.
- target (T.Tensor) – corresponds to a vector that gives the correct label for each example. Labels < 0 are ignored (e.g. can be used for label propagation).
- margin (float or None) –
- scale_correction (float or None) – Downweights absolute deviations for large target scale. The value specifies the target value at which the square deviation has half weight compared to target=0 If the target is twice as large as this value the downweight is 1/3 and so on. Note: the smaller this value the stronger the effect. No effect would be +inf
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
class
elektronn2.neuromancer.loss.AbsLoss(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.loss.SquaredLossAbsLoss node.
Parameters: - pred (Node) – Prediction node.
- target (T.Tensor) – corresponds to a vector that gives the correct label for each example. Labels < 0 are ignored (e.g. can be used for label propagation).
- margin (float or None) –
- scale_correction (float or None) – Boosts loss for large target values: if target=1 the error is multiplied by this value (and linearly for other targets)
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
class
elektronn2.neuromancer.loss.Softmax(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeSoftmax node.
Parameters: - parent (Node) – Input node.
- n_class –
- n_indep –
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
class
elektronn2.neuromancer.loss.MultinoulliNLL(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeReturns the symbolic mean and instance-wise negative log-likelihood of the prediction of this model under a given target distribution.
Parameters: - pred (Node) – Prediction node.
- target (T.Tensor) – corresponds to a vector that gives the correct label for each example. Labels < 0 are ignored (e.g. can be used for label propagation).
- target_is_sparse (bool) – If the target is sparse.
- class_weights (T.Tensor) – weight vector of float32 of length
n_lab. Values:1.0(default),w < 1.0(less important),w > 1.0(more important class). - example_weights (T.Tensor) – weight vector of float32 of shape
(bs, z, x, y)that can give the individual examples (i.e. labels for output pixels) different weights. Values:1.0(default),w < 1.0(less important),w > 1.0(more important example). Note: if this is not normalised/bounded it may result in a effectively modified learning rate! - following refers to lazy labels, the masks are always on a per patch basis, depending on the (The) –
- cube of the patch. The masks are properties of the individual image cubes and must be loaded (origin) –
- CNNData. (into) –
- mask_class_labeled (T.Tensor) – shape = (batchsize, num_classes).
Binary masks indicating whether a class is properly labeled in
y. If a classkis (in general) present in the image patches andmask_class_labeled[k]==1, then the labels must obeyy==kfor all pixels where the class is present. If a classkis present in the image, but was not labeled (-> cheaper labels), setmask_class_labeled[k]=0. Then all pixels for which they==kwill be ignored. Alternative: sety=-1to ignore those pixels. Limit case:mask_class_labeled[:]==1will result in the ordinary NLL. - mask_class_not_present (T.Tensor) – shape = (batchsize, num_classes).
Binary mask indicating whether a class is present in the image patches.
mask_class_not_present[k]==1means that the image does not contain examples of classk. Then for all pixels in the patch, classkpredictive probabilities are trained towards0. Limit case:mask_class_not_present[:]==0will result in the ordinary NLL. - name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
Examples
- A cube contains no class
k. Instead of labelling the remaining classes they can be marked as unlabelled by the first mask (mask_class_labeled[:]==0, whethermask_class_labeled[k]is0or1is actually indifferent because the labels should not bey==kanyway in this case). Additionallymask_class_not_present[k]==1(otherwise0) to suppress predictions ofkin in this patch. The actual value of the labels is indifferent, it can either be-1or it could be the background class, if the background is marked as unlabelled (i.e. then those labels are ignored). - Only part of the cube is densely labelled. Set
mask_class_labeled[:]=1for all classes, but set the label values in the unlabelled part to-1to ignore this part. - Only a particular class
kis labelled in the cube. Either set all other label pixels to-1or the corresponding flags inmask_class_labeledfor the unlabelled classes.
Note
Using
-1labels or telling that a class is not labelled, is somewhat redundant and just supported for convenience.
-
class
elektronn2.neuromancer.loss.MalisNLL(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeMalis NLL node. (See https://github.com/TuragaLab/malis)
Parameters: - pred (Node) – Prediction node.
- aff_gt (T.Tensor) –
- seg_gt (T.Tensor) –
- nhood (np.ndarray) –
- unrestrict_neg (bool) –
- class_weights (T.Tensor) – weight vector of float32 of length
n_lab. Values:1.0(default),w < 1.0(less important),w > 1.0(more important class). - example_weights (T.Tensor) – weight vector of float32 of shape
(bs, z, x, y)that can give the individual examples (i.e. labels for output pixels) different weights. Values:1.0(default),w < 1.0(less important),w > 1.0(more important example). Note: if this is not normalised/bounded it may result in a effectively modified learning rate! - name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
elektronn2.neuromancer.loss.Errors(pred, target, target_is_sparse=False, n_class='auto', n_indep='auto', name='errors', print_repr=True)[source]¶
-
class
elektronn2.neuromancer.loss.BetaNLL(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeSimilar to BinaryNLL loss but “modulated” in scale by the variance.
Parameters: Computes element-wise:
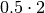
-
elektronn2.neuromancer.loss.SobelizedLoss(pred, target, loss_type='abs', loss_kwargs=None)[source]¶ SobelizedLoss node.
Parameters: - pred (Node) – Prediction node.
- target (T.Tensor) – corresponds to a vector that gives the correct label for each example. Labels < 0 are ignored (e.g. can be used for label propagation).
- loss_type (str) – Only “abs” is supported.
- loss_kwargs (dict) – kwargs for the AbsLoss constructor.
Returns: The loss node.
Return type:
-
class
elektronn2.neuromancer.loss.BlockedMultinoulliNLL(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeReturns the symbolic mean and instance-wise negative log-likelihood of the prediction of this model under a given target distribution.
Parameters: - pred (Node) – Prediction node.
- target (T.Tensor) – corresponds to a vector that gives the correct label for each example. Labels < 0 are ignored (e.g. can be used for label propagation).
- blocking_factor (float) – Blocking factor.
- target_is_sparse (bool) – If the target is sparse.
- class_weights (T.Tensor) – weight vector of float32 of length
n_lab. Values:1.0(default),w < 1.0(less important),w > 1.0(more important class). - example_weights (T.Tensor) – weight vector of float32 of shape
(bs, z, x, y)that can give the individual examples (i.e. labels for output pixels) different weights. Values:1.0(default),w < 1.0(less important),w > 1.0(more important example). Note: if this is not normalised/bounded it may result in a effectively modified learning rate! - following refers to lazy labels, the masks are always on a per patch basis, depending on the (The) –
- cube of the patch. The masks are properties of the individual image cubes and must be loaded (origin) –
- CNNData. (into) –
- mask_class_labeled (T.Tensor) – shape = (batchsize, num_classes).
Binary masks indicating whether a class is properly labeled in
y. If a classkis (in general) present in the image patches andmask_class_labeled[k]==1, then the labels must obeyy==kfor all pixels where the class is present. If a classkis present in the image, but was not labeled (-> cheaper labels), setmask_class_labeled[k]=0. Then all pixels for which they==kwill be ignored. Alternative: sety=-1to ignore those pixels. Limit case:mask_class_labeled[:]==1will result in the ordinary NLL. - mask_class_not_present (T.Tensor) – shape = (batchsize, num_classes).
Binary mask indicating whether a class is present in the image patches.
mask_class_not_present[k]==1means that the image does not contain examples of classk. Then for all pixels in the patch, classkpredictive probabilities are trained towards0. Limit case:mask_class_not_present[:]==0will result in the ordinary NLL. - name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
Examples
- A cube contains no class
k. Instead of labelling the remaining classes they can be marked as unlabelled by the first mask (mask_class_labeled[:]==0, whethermask_class_labeled[k]is0or1is actually indifferent because the labels should not bey==kanyway in this case). Additionallymask_class_not_present[k]==1(otherwise0) to suppress predictions ofkin in this patch. The actual value of the labels is indifferent, it can either be-1or it could be the background class, if the background is marked as unlabelled (i.e. then those labels are ignored). - Only part of the cube is densely labelled. Set
mask_class_labeled[:]=1for all classes, but set the label values in the unlabelled part to-1to ignore this part. - Only a particular class
kis labelled in the cube. Either set all other label pixels to-1or the corresponding flags inmask_class_labeledfor the unlabelled classes.
Note
Using
-1labels or telling that a class is not labelled, is somewhat redundant and just supported for convenience.
-
class
elektronn2.neuromancer.loss.OneHot(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeOnehot node.
Parameters: - target (T.Tensor) – Target tensor.
- n_class (int) –
- axis –
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
class
elektronn2.neuromancer.loss.EuclideanDistance(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeEuclidean distance node.
Parameters: - pred (Node) – Prediction node.
- target (T.Tensor) – corresponds to a vector that gives the correct label for each example. Labels < 0 are ignored (e.g. can be used for label propagation).
- margin (float/None) –
- scale_correction (float/None) – Downweights absolute deviations for large target scale. The value specifies the target value at which the square deviation has half weight compared to target=0 If the target is twice as large as this value the downweight is 1/3 and so on. Note: the smaller this value the stronger the effect. No effect would be +inf
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
class
elektronn2.neuromancer.loss.RampLoss(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeRampLoss node.
Parameters: - pred (Node) – Prediction node.
- target (T.Tensor) – corresponds to a vector that gives the correct label for each example. Labels < 0 are ignored (e.g. can be used for label propagation).
- margin (float/None) –
- scale_correction (float/None) – downweights absolute deviations for large target scale. The value specifies the target value at which the square deviation has half weight compared to target=0 If the target is twice as large as this value the downweight is 1/3 and so on. Note: the smaller this value the stronger the effect. No effect would be +inf
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
elektronn2.neuromancer.model module¶
-
class
elektronn2.neuromancer.model.Model(name='')[source]¶ Bases:
elektronn2.neuromancer.graphmanager.GraphManagerRepresents a neural network model and its current training state.
The Model is defined and checked by running Model.designate_nodes() with appropriate Nodes as arguments (see example in examples/numa_mnist.py).
Permanently saving a Model with its respective training state is possible with the Model.save() function. Loading a Model from a file is done by elektronn2.neuromancer.model.modelload().
During training of a neural network, you can access the current Model via the interactive training shell as the variable “model” (see elektronn2.training.trainutils.user_input()). There are several statistics and hyperparameters of the Model that you can inspect and set directly in the shell, e.g. entering >>> model.lr = 1e-3 and exiting the prompt again effectively sets the learning rate to 1e-3 for further training. (This can also be done with the shortcut “setlr 1e-3”.)
-
batch_normalisation_active¶ Check if batch normalisation is active in any Node in the Model.
-
debug_output_names¶ If debug_outputs is set, a list of all debug output names is returned.
-
designate_nodes(input_node='input', target_node=None, loss_node=None, prediction_node=None, prediction_ext=None, error_node=None, debug_outputs=None)[source]¶ Register input, target and other special Nodes in the Model.
Most of the Model’s attributes are derived from the Nodes that are given as arguments here.
-
dropout_rates¶ Get dropout rates.
-
get_param_values(skip_const=False, as_list=False)[source]¶ Only use this to save/load parameters!
Returns a dict of mapping the values of the params (such that they can be saved to disk) :param skip_const: whether to exclude constant parameters
-
gradnet_rates¶ Get gradnet rates.
Description: https://arxiv.org/abs/1511.06827
-
loss_input_shapes¶ Get shape(s) of loss nodes’ input node(s).
The return value is either a shape (if one input) or a list of shapes (if multiple inputs).
-
loss_smooth¶ Get average loss during the last training steps.
The average is calculated over the last n steps, where n is defined by the config variable time_per_step_smoothing_length (default: 50).
-
lr¶ Get learning rate.
-
measure_exectimes(n_samples=5, n_warmup=4, print_info=True)[source]¶ Return an OrderedDict that maps node names to their estimated execution times in milliseconds.
Parameters are the same as in elektronn2.neuromancer.node_basic.Node.measure_exectime()
-
mixing¶ Get mixing weights.
-
mom¶ Get momentum.
-
prediction_feature_names¶ If a prediction node is set, return its feature names.
-
save(file_name)[source]¶ Save a Model (including its training state) to a pickle file. :param file_name: File name to save the Model in.
-
set_param_values(value_dict, skip_const=False)[source]¶ Only use this to save/load parameters!
Sets new values for non constant parameters :param value_dict: dict mapping values by parameter name / or file name thereof :param skip_const: if dict also maps values for constants, these can be skipped, otherwise an exception is raised.
-
time_per_step¶ Get average run time per training step.
The average is calculated over the last n steps, where n is defined by the config variable time_per_step_smoothing_length (default: 50).
-
trainingstep(*args, **kwargs)[source]¶ Perform one optimiser iteration. Optimisers can be chosen by the kwarg
optimiser.Signature:
trainingstep(data, target(, *aux)(, **kwargs**))Parameters: - *args –
- data: floatX array
- input [bs, ch (, z, y, x)]
- targets: int16 array
- [bs,((z,)y,x)] (optional)
- (optional) other inputs: np.ndarray
- depending in model
- **kwargs –
- optimiser: str
- Name of the chosen optimiser class in
elektronn2.neuromancer.optimiser
- update_loss: Bool
- determine current loss after update step
(e.g. needed for queue, but
get_losscan also be called explicitly)
Returns: - loss (floatX) – Loss (nll, squared error etc…)
- t (float) – Time spent on the GPU per step
- *args –
-
wd¶ Get weight decay.
-
-
elektronn2.neuromancer.model.modelload(file_name, override_mfp_to_active=False, imposed_patch_size=None, imposed_batch_size=None, name=None, **model_load_kwargs)[source]¶ Load a Model from a pickle file (created by Model.save()).
model_load_kwargs: remove_bn, make_weights_constant (True/False)
-
elektronn2.neuromancer.model.kernel_lists_from_node_descr(model_descr)[source]¶ Extract the tuple (filter_shapes, pool_shapes, mfp) from a model description.
Parameters: model_descr – Model description OrderedDict. Returns: Tuple (filter_shapes, pool_shapes, mfp).
-
elektronn2.neuromancer.model.params_from_model_file(file_name)[source]¶ Load parameters from a model file.
Parameters: file_name – File name of the pickled Model. Returns: OrderedDict of model parameters.
-
elektronn2.neuromancer.model.rebuild_model(model, override_mfp_to_active=False, imposed_patch_size=None, name=None, **model_load_kwargs)[source]¶ Rebuild a Model by saving it to a file and reloading it from there.
Parameters: - model – Model object.
- override_mfp_to_active – (See elektronn2.neuromancer.model.modelload()).
- imposed_patch_size – (See elektronn2.neuromancer.model.modelload()).
- name – New model name.
- model_load_kwargs – Additional kwargs for restoring Model (see elektronn2.neuromancer.graphmanager.GraphManager.restore()).
Returns: Rebuilt Model.
-
elektronn2.neuromancer.model.simple_cnn(batch_size, n_ch, n_lab, desired_input, filters, nof_filters, activation_func, pools, mfp=False, tags=None, name=None)[source]¶ Create a simple Model of a convolutional neural network. :param batch_size: Batch size (how many data samples are used in one
update step).Parameters: - n_ch – Number of channels.
- n_lab – Number of distinct labels (classes).
- desired_input – Desired input image size. (Must be smaller than the size of the training images).
- filters – List of filter sizes in each layer.
- nof_filters – List of number of filters for each layer.
- activation_func – Activation function.
- pools – List of maxpooling factors for each layer.
- mfp – List of bools that tell if max fragment pooling should be used in each layer (only intended for prediction).
- tags – Tuple of tags for Input node (see docs of elektronn2.neuromancer.node_basic.Input).
- name – Name of the model.
Returns: Network Model.
elektronn2.neuromancer.neural module¶
-
class
elektronn2.neuromancer.neural.Perceptron(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.neural.NeuralLayerPerceptron Layer.
Parameters: - parent (Node or list of Node) – The input node(s).
- n_f (int) – Number of filters (nodes) in layer.
- activation_func (str) – Activation function name.
- flatten (bool) –
- batch_normalisation (str or None) – Batch normalisation mode. Can be False (inactive), “train” or “fadeout”.
- dropout_rate (float) – Dropout rate (probability that a node drops out in a training step).
- name (str) – Perceptron name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
- w (np.ndarray or T.TensorVariable) – Weight matrix. If this is a np.ndarray, its values are used to initialise a shared variable for this layer. If it is a T.TensorVariable, it is directly used (weight sharing with the layer which this variable comes from).
- b (np.ndarray or T.TensorVariable) – Bias vector. If this is a np.ndarray, its values are used to initialise a shared variable for this layer. If it is a T.TensorVariable, it is directly used (weight sharing with the layer which this variable comes from).
- gamma – (For batch normalisation) Initialises gamma parameter.
- mean – (For batch normalisation) Initialises mean parameter.
- std – (For batch normalisation) Initialises std parameter.
- gradnet_mode –
-
make_dual(parent, share_w=False, **kwargs)[source]¶ Create the inverse of this
Perceptron.Most options are the same as for the layer itself. If
kwargsare not specified, the values of the primal layers are re-used and new parameters are created.Parameters: - parent (Node) – The input node.
- share_w (bool) – If the weights (
w) should be shared from the primal layer. - kwargs (dict) – kwargs that are passed through to the constructor of the inverted
Perceptron (see signature of
Perceptron).n_fis copied from the existing node on whichmake_dualis called. Every other parameter can be changed from the originalPerceptron’s defaults by specifying it inkwargs.
Returns: The inverted perceptron layer.
Return type:
-
class
elektronn2.neuromancer.neural.Conv(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.neural.PerceptronConvolutional layer with subsequent pooling.
Examples
Examples for constructing convolutional neural networks can be found in examples/neuro3d.py and examples/mnist.py.
Parameters: - parent (Node) – The input node.
- n_f (int) – Number of features.
- filter_shape (tuple) – Shape of the convolution filter kernels.
- pool_shape (tuple) – Shape of max-pooling to be applied after the convolution.
None(default) disables pooling along all axes. - conv_mode (str) –
Possible values: * “valid”: Only apply filter to complete patches of the image.
Generates output of shape: image_shape -filter_shape + 1.- ”full”: Zero-pads image to multiple of filter shape to generate output of shape: image_shape + filter_shape - 1.
- ”same”: Zero-pads input image so that the output shape is equal to the input shape (Only supported for odd filter sizes).
- activation_func (str) – Activation function name.
- mfp (bool) – Whether to apply Max-Fragment-Pooling in this Layer.
- batch_normalisation (str or False) – Batch normalisation mode. Can be False (inactive), “train” or “fadeout”.
- dropout_rate (float) – Dropout rate (probability that a node drops out in a training step).
- name (str) – Layer name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
- w (np.ndarray or T.TensorVariable) – Weight matrix. If this is a np.ndarray, its values are used to initialise a shared variable for this layer. If it is a T.TensorVariable, it is directly used (weight sharing with the layer which this variable comes from).
- b (np.ndarray or T.TensorVariable) – Bias vector. If this is a np.ndarray, its values are used to initialise a shared variable for this layer. If it is a T.TensorVariable, it is directly used (weight sharing with the layer which this variable comes from).
- gamma – (For batch normalisation) Initialises gamma parameter.
- mean – (For batch normalisation) Initialises mean parameter.
- std – (For batch normalisation) Initialises std parameter.
- gradnet_mode –
- invalidate_fov (bool) – Overrides the computed
fovwith an invalid value to force later recalculation (experimental).
-
make_dual(parent, share_w=False, mfp=False, **kwargs)[source]¶ Create the inverse (
UpConv) of thisConvnode.Most options are the same as for the layer itself. If
kwargsare not specified, the values of the primal layers are re-used and new parameters are created.Parameters: - parent (Node) – The input node.
- share_w (bool) – If the weights (
w) should be shared from the primal layer. - mfp (bool) – If max-fragment-pooling is used.
- kwargs (dict) – kwargs that are passed through to the new
UpConvnode (see signature ofUpConv).n_fandpool_shapeare copied from the existing node on whichmake_dualis called. Every other parameter can be changed from the originalConv’s defaults by specifying it inkwargs.
Returns: The inverted conv layer (as an
UpConvnode).Return type:
-
class
elektronn2.neuromancer.neural.UpConv(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.neural.ConvUpconvolution layer. Also known as transposed convolution.
E.g. pooling + upconv with pool_shape = 3:
x x x x x x x x x before pooling (not in this layer) \|/ \|/ \|/ pooling (not in this layer) x x x input to this layer 0 0 x 0 0 x 0 0 x 0 0 unpooling + padding (done in this layer) /|\ /|\ /|\ conv on unpooled (done in this layer) y y y y y y y y y result of this layer
Parameters: - parent (Node) – The input node.
- n_f (int) – Number of filters (nodes) in layer.
- pool_shape (tuple) – Size of the UpConvolution.
- activation_func (str) – Activation function name.
- identity_init (bool) – Initialise weights to result in pixel repetition upsampling
- batch_normalisation (str or False) – Batch normalisation mode. Can be False (inactive), “train” or “fadeout”.
- dropout_rate (float) – Dropout rate (probability that a node drops out in a training step).
- name (str) – Layer name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
- w (np.ndarray or T.TensorVariable) – Weight matrix. If this is a np.ndarray, its values are used to initialise a shared variable for this layer. If it is a T.TensorVariable, it is directly used (weight sharing with the layer which this variable comes from).
- b (np.ndarray or T.TensorVariable) – Bias vector. If this is a np.ndarray, its values are used to initialise a shared variable for this layer. If it is a T.TensorVariable, it is directly used (weight sharing with the layer which this variable comes from).
- gamma – (For batch normalisation) Initialises gamma parameter.
- mean – (For batch normalisation) Initialises mean parameter.
- std – (For batch normalisation) Initialises std parameter.
- gradnet_mode –
-
make_dual(*args, **kwargs)[source]¶ Create the inverse (
UpConv) of thisConvnode.Most options are the same as for the layer itself. If
kwargsare not specified, the values of the primal layers are re-used and new parameters are created.Parameters: - parent (Node) – The input node.
- share_w (bool) – If the weights (
w) should be shared from the primal layer. - mfp (bool) – If max-fragment-pooling is used.
- kwargs (dict) – kwargs that are passed through to the new
UpConvnode (see signature ofUpConv).n_fandpool_shapeare copied from the existing node on whichmake_dualis called. Every other parameter can be changed from the originalConv’s defaults by specifying it inkwargs.
Returns: - The inverted conv layer (as an
UpConvnode). NOTE: docstring was inherited
Return type:
-
class
elektronn2.neuromancer.neural.Crop(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeThis node type crops the output of its parent.
Parameters: - parent (Node) – The input node whose output should be cropped.
- crop (tuple or list of ints) – Crop each spatial axis from either side by this number.
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
class
elektronn2.neuromancer.neural.LSTM(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.neural.NeuralLayerLong short term memory layer.
Using an implementation without peepholes in f, i, o, i.e. weights cell state is not taken into account for weights. See http://colah.github.io/posts/2015-08-Understanding-LSTMs/.
Parameters: - parent (Node) – The input node.
- memory_states (Node) – Concatenated (initial) feed-back and cell state (one Node!).
- n_f (int) – Number of features.
- activation_func (str) – Activation function name.
- flatten –
- batch_normalisation (str or None) – Batch normalisation mode. Can be False (inactive), “train” or “fadeout”.
- dropout_rate (float) – Dropout rate (probability that a node drops out in a training step).
- name (str) – Layer name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
- w (np.ndarray or T.TensorVariable) – Weight matrix. If this is a np.ndarray, its values are used to initialise a shared variable for this layer. If it is a T.TensorVariable, it is directly used (weight sharing with the layer which this variable comes from).
- b (np.ndarray or T.TensorVariable) – Bias vector. If this is a np.ndarray, its values are used to initialise a shared variable for this layer. If it is a T.TensorVariable, it is directly used (weight sharing with the layer which this variable comes from).
- gamma – (For batch normalisation) Initialises gamma parameter.
- mean – (For batch normalisation) Initialises mean parameter.
- std – (For batch normalisation) Initialises std parameter.
- gradnet_mode –
-
class
elektronn2.neuromancer.neural.Pool(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodePooling layer.
Reduces the count of training parameters by reducing the spatial size of its input by the factors given in
pool_shape.Pooling modes other than max-pooling can only be selected if cuDNN is available.
Parameters: - parent (Node) – The input node.
- pool_shape (tuple) – Tuple of pooling factors (per dimension) by which the input is downsampled.
- stride (tuple) – Stride sizes (per dimension).
- mfp (bool) – If max-fragment-pooling should be used.
- mode (str) –
(only if cuDNN is available) Mode can be any of the modes supported by Theano’s dnn_pool(): (‘max’, ‘average_inc_pad’, ‘average_exc_pad’, ‘sum’).
’max’ (default): max-pooling ‘average’ or ‘average_inc_pad’: average-pooling ‘sum’: sum-pooling
- name (str) – Name of the pooling layer.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
elektronn2.neuromancer.neural.Dot¶ alias of
Perceptron
-
class
elektronn2.neuromancer.neural.FaithlessMerge(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeFaithlessMerge node.
Parameters: - hard_features (Node) –
- easy_features (Node) –
- axis –
- failing_prob (float) – The higher the more often merge is unreliable
- hardeasy_ratio (float) – The higher the more often the harder features fail instead of the easy ones
- name (str) –
Name of the pooling layer. print_repr: bool
Whether to print the node representation upon initialisation.
-
class
elektronn2.neuromancer.neural.GRU(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.neural.NeuralLayerGated Recurrent Unit Layer.
Parameters: - parent (Node) – The input node.
- memory_state (Node) – Memory node.
- n_f (int) – Number of features.
- activation_func (str) – Activation function name.
- flatten (bool) – (Unsupported).
- batch_normalisation (str or None) – Batch normalisation mode. Can be False (inactive), “train” or “fadeout”.
- dropout_rate (float) – Dropout rate (probability that a node drops out in a training step).
- name (str) – Layer name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
- w (np.ndarray or T.TensorVariable) – (Unsupported). Weight matrix. If this is a np.ndarray, its values are used to initialise a shared variable for this layer. If it is a T.TensorVariable, it is directly used (weight sharing with the layer which this variable comes from).
- b (np.ndarray or T.TensorVariable) – (Unsupported). Bias vector. If this is a np.ndarray, its values are used to initialise a shared variable for this layer. If it is a T.TensorVariable, it is directly used (weight sharing with the layer which this variable comes from).
- gamma – (For batch normalisation) Initialises gamma parameter.
- mean – (For batch normalisation) Initialises mean parameter.
- std – (For batch normalisation) Initialises std parameter.
- gradnet_mode –
-
class
elektronn2.neuromancer.neural.LRN(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeLRN (Local Response Normalization) layer.
Parameters: - parent (Node) – The input node.
- filter_shape (tuple) –
- mode (str) – Can be “spatial” or “channel”.
- alpha (float) –
- k (float) –
- beta (float) –
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
elektronn2.neuromancer.neural.AutoMerge(parent1, parent2, upconv_n_f=None, merge_mode='concat', disable_upconv=False, upconv_kwargs=None, name='merge', print_repr=True)[source]¶ Merges two network branches by automatic cropping and upconvolutions.
(Wrapper for
UpConv,Crop,ConcatandAdd.)Tries to automatically align and merge a high-res and a low-res (convolution) output of two branches of a CNN by applying UpConv and Crop to make their shapes and strides compatible. UpConv is used if the low-res parent’s strides are at least twice as large as the strides of the high-res parent in any dimension.
The parents are automatically identified as high-res and low-res by their strides. If both parents have the same strides, the concept of high-res and low-res is ignored and this function just crops the larger parent’s output until the parents’ spatial shapes match and then merges them.
This function can be used to simplify creation of e.g. architectures similar to U-Net (see https://arxiv.org/abs/1505.04597) or skip-connections.
If a ValueError that the shapes cannot be aligned is thrown, you can try changing the filter shapes and pooling factors of the (grand-)parent nodes or add/remove Convolutions and Crops in the preceding branches until the error disappears (of course you should try to keep those changes as minimal as possible).
(This function is an alias for UpConvMerge.)
Parameters: - parent1 (Node) – First parent to be merged.
- parent2 (Node) – Second parent to be merged.
- upconv_n_f (int) – Number of filters for the aligning
UpConvfor the low-res parent. - merge_mode (str) – How the merging should be performed. Available options:
‘concat’ (default): Merge with a
Concatnode. ‘add’: Merge with anAddnode. - disable_upconv (bool) – If
True, no automatic upconvolutions are performed to match strides. - upconv_kwargs (dict) – Additional keyword arguments that are passed to the
UpConvconstructor if upconvolution is applied. - name (str) – Name of the final merge node.
- print_repr (bool) – Whether to print the node representation upon initialisation.
Returns: ConcatorAddnode (depending onmerge_mode) that merges the aligned high-res and low-res outputs.Return type:
-
elektronn2.neuromancer.neural.UpConvMerge(parent1, parent2, upconv_n_f=None, merge_mode='concat', disable_upconv=False, upconv_kwargs=None, name='merge', print_repr=True)¶ Merges two network branches by automatic cropping and upconvolutions.
(Wrapper for
UpConv,Crop,ConcatandAdd.)Tries to automatically align and merge a high-res and a low-res (convolution) output of two branches of a CNN by applying UpConv and Crop to make their shapes and strides compatible. UpConv is used if the low-res parent’s strides are at least twice as large as the strides of the high-res parent in any dimension.
The parents are automatically identified as high-res and low-res by their strides. If both parents have the same strides, the concept of high-res and low-res is ignored and this function just crops the larger parent’s output until the parents’ spatial shapes match and then merges them.
This function can be used to simplify creation of e.g. architectures similar to U-Net (see https://arxiv.org/abs/1505.04597) or skip-connections.
If a ValueError that the shapes cannot be aligned is thrown, you can try changing the filter shapes and pooling factors of the (grand-)parent nodes or add/remove Convolutions and Crops in the preceding branches until the error disappears (of course you should try to keep those changes as minimal as possible).
(This function is an alias for UpConvMerge.)
Parameters: - parent1 (Node) – First parent to be merged.
- parent2 (Node) – Second parent to be merged.
- upconv_n_f (int) – Number of filters for the aligning
UpConvfor the low-res parent. - merge_mode (str) – How the merging should be performed. Available options:
‘concat’ (default): Merge with a
Concatnode. ‘add’: Merge with anAddnode. - disable_upconv (bool) – If
True, no automatic upconvolutions are performed to match strides. - upconv_kwargs (dict) – Additional keyword arguments that are passed to the
UpConvconstructor if upconvolution is applied. - name (str) – Name of the final merge node.
- print_repr (bool) – Whether to print the node representation upon initialisation.
Returns: ConcatorAddnode (depending onmerge_mode) that merges the aligned high-res and low-res outputs.Return type:
-
class
elektronn2.neuromancer.neural.Pad(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodePads the spatial axes of its parent’s output.
Parameters: - parent (Node) – The input node whose output should be padded.
- pad (tuple or list of ints) – The padding length from either side for each spatial axis
- value (float) – Value of the padding elements (default: 0.0)
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
elektronn2.neuromancer.node_basic module¶
-
class
elektronn2.neuromancer.node_basic.Node(parent, name='', print_repr=False)[source]¶ Bases:
objectBasic node class. All neural network nodes should inherit from
Node.Parameters: Models are built from the interplay of Nodes to form a (directed, acyclic) computational graph.
The ELEKTRONN2 framework can be seen as an intelligent abstraction level that hides the raw theano-graph and manages the involved symbolic variables. The overall goal is the intuitive, flexible and easy creation of complicated graphs.
A
Nodehas one or several inputs, calledparent, (unless it is a source, i.e. a node where external data is feed into the graph). The inputs are node objects themselves.Layers automatically keep track of their previous inputs, parameters, computational cost etc. This allows to compile the theano-functions without manually specifying the inputs, outputs and parameters. In the most simple case, any node, which might be part of a more complicated graph, can be called as a function (passing suitable numpy arrays):
>>> import elektronn2.neuromancer.utils >>> inp = neuromancer.Input((batch_size, in_dim)) >>> test_data = elektronn2.neuromancer.utils.as_floatX(np.random.rand(batch_size, in_dim)) >>> out = inp(test_data) >>> np.allclose(out, test_data) True
At the first time the theano function is compiled and cached for re-use in future calls.
Several properties (with respect to the sub-graph the node depends on, or only from the of the node itself) These can also be looked up externally (e.g. required sources, parameter count, computational count).
The theano variable that represents the output of a node is kept in the attribute
output. Subsequent Nodes must use this attribute of their inputs to perform their calculation and write the result to their own output (this happens in the method_calc_output, which is hidden because it must be called only internally at initialisation).A divergence in the computational graph is created by passing the parent to several children as input:
>>> inp = neuromancer.Input((1,10), name='Input_1') >>> node1 = neuromancer.ApplyFunc(inp, func1) >>> node2 = neuromancer.ApplyFunc(inp, func2)
A convergence in the graph is created by passing several inputs to a node that performs a reduction:
>>> out = neuromancer.Concat([node1, node2])
Although the node “out” has two immediate inputs, it is detected that the required sources is only a single object:
>>> print(out.input_nodes) Input_1
Computations that result in more than a single output for a Node must be broken apart using divergence and individual nodes for the several outputs. Alternatively the function
splitcan be used to create two dummy nodes of the output of a previous Node by splitting along the specified axis. Note that possible redundant computations in Nodes are most likely eliminated by the theano graph optimiser.Instructions for subclassing:
- Overriding
__init__: - At the very first the base class’ initialiser must be called, which just
assigns the names and emtpy default values for attributes.
Then node specific initialisations are made e.g. initialisation of shared
parameters / weights.
Finally the
_finialise_initmethod of the base class is automatically called: This evokes the execution of the methods:_make_output,_calc_shapeandself._calc_comp_cost. Each of those updates the corresponding attributes. NOTE: if a Node (except for the baseNode) is subclassed and the derived calls__init__of the base Node, this will also call_finialise_initexactly right the call to the superclass’__init__.
For the graph serialisation and restoration to work, the following conditions must additionally be met:
- The name of of a node’s trainable parameter in the parameter dict must
be the same as the (optional) keyword used to initialise this parameter
in
__init__; moreover, parameters must not be initialised/shared from positional arguments. - When serialising only the current state of parameters is kept, parameter value arrays given for initialisation are never kept.
Depending on the purpose of the node, the latter methods and others (e.g.
__repr__) must be overridden. The default behaviour of the base class is: output = input, outputs shape = input shape, computational cost = tensor size (!) …-
all_children¶
-
all_computational_cost¶
-
all_extra_updates¶ List of the parameters updates of all parent nodes. They are tuples.
-
all_nontrainable_params¶ Dict of the trainable parameters (weights) of all parent nodes. They are theano shared variables.
-
all_params¶ Dict of the all parameters of all parent nodes. They are theano variable
-
all_params_count¶ Count of all trainable parameters in the entire sub-graph used to compute the output of this node
-
all_parents¶ List all nodes that are involved in the computation of the output of this node (incl.
self). The list contains no duplicates. The return is a dict, the keys of which are the layers, the values are just allTrue
-
all_trainable_params¶ Dict of the trainable parameters (weights) of all parent nodes. They are theano shared variables.
-
feature_names¶
-
get_param_values(skip_const=False)[source]¶ Returns a dict that maps the values of the params. (such that they can be saved to disk)
Parameters: skip_const (bool) – whether to exclude constant parameters. Returns: Dict that maps the values of the params. Return type: dict
-
input_nodes¶ Contains the all parent nodes that are sources, i.e. inputs that are required to compute the result of this node.
-
input_tensors¶ The same as
input_nodesbut contains the theano tensor variables instead of the node objects. May be used as input to compile theano functions.
-
last_exec_time¶ Last function execution time in seconds.
-
local_exec_time¶
-
measure_exectime(n_samples=5, n_warmup=4, print_info=True, local=True, nonegative=True)[source]¶ Measure how much time the node needs for its calculation (in milliseconds).
Parameters: - n_samples (int) – Number of independent measurements of which the median is taken.
- n_warmup (int) – Number of warm-up runs before each measurement (not taken into account for median calculation).
- print_info (bool) – If True, print detailed info about measurements while running.
- local (bool) – Only compute exec time for this node by subtracting its parents’ times.
- nonegative (bool) – Do not return exec times smaller than zero.
Returns: median of execution time measurements.
Return type: np.float
-
param_count¶ Count of trainable parameters in this node
-
plot_theano_graph(outfile=None, compiled=True, **kwargs)[source]¶ Plot the execution graph of this Node’s Theano function to a file.
If “outfile” is not specified, the plot is saved in “/tmp/<user>_<name>.png”
Parameters: - outfile (str or None) – File name for saving the plot.
- compiled (bool) – If True, the function is compiled before plotting.
- kwargs – kwargs (plotting options) that get directly passed to theano.printing.pydotprint().
-
predict_dense(raw_img, as_uint8=False, pad_raw=False)[source]¶ Core function that performs the inference
Parameters: - raw_img (np.ndarray) – raw data in the format (ch, (z,) y, x)
- as_uint8 (Bool) – Return class probabilites as uint8 image (scaled between 0 and 255!)
- pad_raw (Bool) – Whether to apply padding (by mirroring) to the raw input image in order to get predictions on the full image domain.
Returns: Predictions.
Return type: np.ndarray
-
set_param_values(value_dict, skip_const=False)[source]¶ Sets new values for non constant parameters.
Parameters: - value_dict (dict) – A dict that maps values by parameter name.
- skip_const (bool) – if dict also maps values for constants, these can be skipped, otherwise an exception is raised.
-
test_run(on_shape_mismatch='warn', debug_outputs=False)[source]¶ Test execution of this node with random (but correctly shaped) data.
Parameters: on_shape_mismatch (str) – If this is “warn”, a warning is emitted if there is a mismatch between expected and calculated output shapes. Returns: Return type: Debug output of the Theano function.
-
total_exec_time¶
- Overriding
-
class
elektronn2.neuromancer.node_basic.Input(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeInput node
Parameters: - shape (list/tuple of int) – shape of input array, unspecified shapes are
None - tags (list/tuple of strings or comma-separated string) – tags indicate which purpose the dimensions of the tensor serve. They are
sometimes used to decide about reshapes. The maximal tensor has tags:
“r, b, f, z, y, x, s” which denote:
* r: perform recurrence along this axis
* b: batch size
* f: features, filters, channels
* z: convolution no. 3 (slower than 1,2)
* y: convolution no. 1
* x: convolution no. 2
* s: samples of the same instance (over which expectations are calculated)
Unused axes are to be removed from this list, but
bandfmust always remain. To avoid bad memory layout, the order must not be changed. For less than 3 convolutions conv1,conv2 are preferred for performance reasons. Note that CNNs can mix nodes with 2d and 3d convolutions as 2d is a special case of 3d with filter size 1 on the respective axis. In this case conv3 should be used for the axis with smallest filter size. - strides –
- fov –
- dtype (str) – corresponding to numpy dtype (e.g., ‘int64’). Default is floatX from theano config
- hardcoded_shape –
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
- shape (list/tuple of int) – shape of input array, unspecified shapes are
-
elektronn2.neuromancer.node_basic.Input_like(ref, dtype=None, name='input', print_repr=True, override_f=False, hardcoded_shape=False)[source]¶
-
class
elektronn2.neuromancer.node_basic.Concat(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeNode to concatenate the inputs. The inputs must have the same shape, except in the dimension corresponding to
axis. This is not checked as shapes might be unspecified prior to compilation!Parameters: - parent_nodes (list of Node) – Inputs to be concatenated.
- axis (int) – Join axis.
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
class
elektronn2.neuromancer.node_basic.ApplyFunc(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeApply function to the input. If the function changes the output shape, this node should not be used.
Parameters: - parent (Node) – Input (single).
- functor (function) – Function that acts on theano variables (e.g.
theano.tensor.tanh). - args (tuple) – Arguments passed to
functorafter the input. - kwargs (dict) – kwargs for
functor.
-
class
elektronn2.neuromancer.node_basic.FromTensor(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeDummy node to be used in the split-function.
Parameters: - tensor (T.Tensor) –
- tensor_shape –
- tensor_parent (T.Tensor) –
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
elektronn2.neuromancer.node_basic.split(node, axis='f', index=None, n_out=None, strip_singleton_dims=False, name='split')[source]¶
-
class
elektronn2.neuromancer.node_basic.GenericInput(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeInput node for arbitrary oject.
Parameters: - name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
-
class
elektronn2.neuromancer.node_basic.ValueNode(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.Node(Optionally) trainable value node
Parameters: - shape (list/tuple of int) – shape of input array, unspecified shapes are
None - tags (list/tuple of strings or comma-separated string) –
tags indicate which purpose the dimensions of the tensor serve. They are sometimes used to decide about reshapes. The maximal tensor has tags: “r, b, f, z, y, x, s” which denote:
- r: perform recurrence along this axis
- b: batch size
- f: features, filters, channels
- z: convolution no. 3 (slower than 1,2)
- y: convolution no. 1
- x: convolution no. 2
- s: samples of the same instance (over which expectations are calculated)
Unused axes are to be removed from this list, but
bandfmust always remain. To avoid bad memory layout, the order must not be changed. For less than 3 convolutions conv1,conv2 are preferred for performance reasons. Note that CNNs can mix nodes with 2d and 3d convolutions as 2d is a special case of 3d with filter size 1 on the respective axis. In this case conv3 should be used for the axis with smallest filter size. - strides –
- fov –
- dtype (str) – corresponding to numpy dtype (e.g., ‘int64’). Default is floatX from theano config
- apply_train (bool) –
- value –
- init_kwargs (dict) –
- name (str) – Node name.
- print_repr (bool) – Whether to print the node representation upon initialisation.
- shape (list/tuple of int) – shape of input array, unspecified shapes are
-
class
elektronn2.neuromancer.node_basic.MultMerge(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeNode to concatenate the inputs. The inputs must have the same shape, except in the dimension corresponding to
axis. This is not checked as shapes might be unspecified prior to compilation!Parameters:
-
class
elektronn2.neuromancer.node_basic.InitialState_like(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeParameters: - parent –
- override_f –
- dtype –
- name –
- print_repr –
- init_kwargs –
-
class
elektronn2.neuromancer.node_basic.Add(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeAdd two nodes using
theano.tensor.add.Parameters:
elektronn2.neuromancer.optimiser module¶
-
class
elektronn2.neuromancer.optimiser.AdaDelta(inputs, loss, grads, params, extra_updates, additional_outputs=None)[source]¶ Bases:
elektronn2.neuromancer.optimiser.OptimiserAdaDelta optimiser (See https://arxiv.org/abs/1212.5701).
Like AdaGrad, but accumulate squared only over window The delta part is some diagonal hessian approximation. Claims to be robust against sudden large gradients because then the denominator explodes, but this explosion is persistent for a while… (and this argumentation is true for any method accumulating squared grads).
-
class
elektronn2.neuromancer.optimiser.AdaGrad(inputs, loss, grads, params, extra_updates, additional_outputs=None)[source]¶ Bases:
elektronn2.neuromancer.optimiser.OptimiserAdaGrad optimiser (See http://jmlr.org/papers/v12/duchi11a.html).
Tries to favor making faster progress on parameters with usually small gradients (but does somehow ignore their actual direction, i.e. a parameter which has a lot of small gradients in the same direction and one that has many small gradients in opposite directions have both a high LR !
-
class
elektronn2.neuromancer.optimiser.Adam(inputs, loss, grads, params, extra_updates, additional_outputs=None)[source]¶ Bases:
elektronn2.neuromancer.optimiser.OptimiserAdam optimiser (See https://arxiv.org/abs/1412.6980v9).
Like AdaGrad with windowed squared_accum and with momentum and a bias for the initial phase (t). The normalisation of Adam and AdaGrad (and RMSProp) does not damp but exaggerates sudden steep gradients (their squared_accum is small and their current grad is large).
-
class
elektronn2.neuromancer.optimiser.Optimiser(inputs, loss, grads, params, additional_outputs)[source]¶ Bases:
objectReturns new shared variables matching the shape of params/gradients
-
global_lr= lr¶
-
global_mom= mom¶
-
global_weight_decay= weight_decay¶
-
class
elektronn2.neuromancer.optimiser.SGD(inputs, loss, grads, params, extra_updates, additional_outputs=None)[source]¶ Bases:
elektronn2.neuromancer.optimiser.OptimiserSGD optimiser (See https://en.wikipedia.org/wiki/Stochastic_gradient_descent).
elektronn2.neuromancer.variables module¶
-
class
elektronn2.neuromancer.variables.VariableParam(value=None, name=None, apply_train=True, apply_reg=True, dtype=None, strict=False, allow_downcast=None, borrow=False, broadcastable=None)[source]¶ Bases:
theano.tensor.sharedvar.TensorSharedVariableExtension of theano
TensorSharedVariable. Additional features are described by the parameters, otherwise identicalParameters: - value –
- name (str) –
- flag (apply_train) – whether to apply regularisation (e.g. L2) on this param
- flag – whether to train this parameter (as opposed to a meta-parameter or a parameter that is kept const. during a training phase)
- dtype –
- strict (bool) –
- allow_downcast (bool) –
- borrow (bool) –
- broadcastable –
-
updates¶
-
class
elektronn2.neuromancer.variables.VariableWeight(shape=None, init_kwargs=None, value=None, name=None, apply_train=True, apply_reg=True, dtype=None, strict=False, allow_downcast=None, borrow=False, broadcastable=None)[source]¶
-
class
elektronn2.neuromancer.variables.ConstantParam(value, name=None, dtype=None, make_singletons_broadcastable=True)[source]¶ Bases:
theano.tensor.var.TensorConstantIdentical to theano
VariableParamexcept that there are two two addition attributesapply_trainand apply_reg`, which are both false. This is just to tell ELEKTRONN2 that this parameter is to be exempted from training. Obviously theset_valuemethod raises an exception because this is a real constant. Constants are faster in the theano graph.-
updates¶
-
elektronn2.neuromancer.various module¶
-
class
elektronn2.neuromancer.various.GaussianRV(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeParameters: - mu (node) – Mean of the Gaussian density
- sig (node) – Sigma of the Gaussian density
- n_samples (int) – Number of samples to be drawn per instance. Special case ‘0’: draw 1 sample but don’t’ increase rank of tensor!
- output is a sample from separable Gaussians of given mean and (The) –
- (but this operation is still differentiable, due to the (sigma) –
- trick") ("re-parameterisation) –
- output dimension mu.ndim+1 because the samples are accumulated along (The) –
- new axis right of 'b' (batch) (a) –
-
elektronn2.neuromancer.various.SkelLoss(pred, loss_kwargs, skel=None, name='skel_loss', print_repr=True)[source]¶
-
class
elektronn2.neuromancer.various.SkelPrior(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.Nodepred must be a vector of shape [(1,b),(3,f)] or [(3,f)] i.e. only batch_size=1 is supported.
Parameters: - pred –
- target_length –
- prior_n –
- prior_posz –
- prior_z –
- prior_xy –
- name –
- print_repr –
-
elektronn2.neuromancer.various.Scan(step_result, in_memory, out_memory=None, in_iterate=None, in_iterate_0=None, n_steps=None, unroll_scan=True, last_only=False, name='scan', print_repr=True)[source]¶ Parameters: - step_result (node/list(nodes)) – nodes that represent results of step function
- in_memory (node/list(nodes)) – nodes that indicate at which place in the computational graph
the memory is feed back into the step function. If
out_memoryis not specified this must contain a node for every node instep_resultbecause then the whole result will be fed back. - out_memory (node/list(nodes)) – (optional) must be subset of
step_resultand of same length asin_memory, tells which nodes of the result are fed back toin_memory. IfNone, all are fed back. - in_iterate (node/list(nodes)) – nodes with a leading
'r'axis to be iterated over (e.g. time series of shape [(30,r),(100,b),(50,f)]). In every step a slice from the first axis is consumed. - in_iterate_0 (node/list(nodes)) – nodes that consume a single slice of the
in_iteratenodes. Part of “the inner function” of the scan loop in contrast toin_iterate - n_steps (int) –
- unroll_scan (bool) –
- last_only (bool) –
- name (str) –
- print_repr (bool) –
Returns: - A node for every node in
step_resultwhich either contains the last - state or the series of states - then it has a leading
'r'axis.
-
elektronn2.neuromancer.various.SkelGetBatch(skel, aux, img_sh, t_img_sh, t_grid_sh, t_node_sh, get_batch_kwargs, scale_strenght=None, name='skel_batch')[source]¶
-
class
elektronn2.neuromancer.various.SkelLossRec(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.Nodepred must be a vector of shape [(1,b),(3,f)] or [(3,f)] i.e. only batch_size=1 is supported.
Parameters: - pred –
- skel –
- loss_kwargs –
- name –
- print_repr –
-
class
elektronn2.neuromancer.various.Reshape(*args, **kwargs)[source]¶ Bases:
elektronn2.neuromancer.node_basic.NodeReshape node.
Parameters: - parent –
- shape –
- tags –
- strides –
- fov –
- name –
- print_repr –